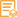Catalog: YM6220
Size
Price
Status
Qty.
200μL
$600.00
In stock
0
100μL
$340.00
In stock
0
40μL
$190.00
In stock
0
Add to cart


Collected


Collect
Main Information
Target
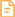
PMS2
Host Species
Mouse
Reactivity
Human, Mouse
Applications
IHC, ELISA
MW
96kD (Calculated)
110kD (Observed)
Conjugate/Modification
Unmodified
Detailed Information
Recommended Dilution Ratio
IHC 1:50-100; ELISA 1:500-5000
Formulation
PBS, 50% glycerol, 0.05% Proclin 300, 0.05%BSA
Specificity
The antibody can specifically recognize human PMS2 protein.
Purification
The antibody was affinity-purified from ascites by affinity-chromatography using specific immunogen.
Storage
-15°C to -25°C/1 year(Do not lower than -25°C)
MW(Calculated)
96kD
MW(Observed)
110kD
Modification
Unmodified
Clonality
Monoclonal
Clone Number
PT2116
Isotype
IgG1,Kappa
Related Products
Antigen&Target Information
Immunogen:
Synthesized peptide derived from human Postmeiotic Segregation Increased 2(PMS2) AA range: 600-700
show all
Specificity:
The antibody can specifically recognize human PMS2 protein.
show all
Gene Name:
PMS2 PMSL2
show all
Protein Name:
Postmeiotic Segregation Increased 2(PMS2)
show all
Other Name:
Mismatch repair endonuclease PMS2 ;
DNA mismatch repair protein PMS2 ;
PMS1 protein homolog 2 ;
DNA mismatch repair protein PMS2 ;
PMS1 protein homolog 2 ;
show all
Background:
The protein encoded by this gene is a key component of the mismatch repair system that functions to correct DNA mismatches and small insertions and deletions that can occur during DNA replication and homologous recombination. This protein forms heterodimers with the gene product of the mutL homolog 1 (MLH1) gene to form the MutL-alpha heterodimer. The MutL-alpha heterodimer possesses an endonucleolytic activity that is activated following recognition of mismatches and insertion/deletion loops by the MutS-alpha and MutS-beta heterodimers, and is necessary for removal of the mismatched DNA. There is a DQHA(X)2E(X)4E motif found at the C-terminus of the protein encoded by this gene that forms part of the active site of the nuclease. Mutations in this gene have been associated with hereditary nonpolyposis colorectal cancer (HNPCC; also known as Lynch syndrome) and Turcot syndrome
show all
Function:
Disease:Defects in PMS2 are a cause of mismatch repair cancer syndrome (MMRCS) [MIM:276300]; also known as Turcot syndrome and brain tumor-polyposis syndrome 1 (BTPS1). MMRCS is an autosomal dominant disorder characterized by malignant tumors of the brain associated with multiple colorectal adenomas. Skin features include sebaceous cysts, hyperpigmented and cafe au lait spots.,Disease:Defects in PMS2 are the cause of hereditary non-polyposis colorectal cancer type 4 (HNPCC4) [MIM:600259]. Mutations in more than one gene locus can be involved alone or in combination in the production of the HNPCC phenotype (also called Lynch syndrome). Most families with clinically recognized HNPCC have mutations in either MLH1 or MSH2 genes. HNPCC is an autosomal, dominantly inherited disease associated with marked increase in cancer susceptibility. It is characterized by a familial predisposition to early onset colorectal carcinoma (CRC) and extra-colonic cancers of the gastrointestinal, urological and female reproductive tracts. HNPCC is reported to be the most common form of inherited colorectal cancer in the Western world, and accounts for 15% of all colon cancers. Cancers in HNPCC originate within benign neoplastic polyps termed adenomas. Clinically, HNPCC is often divided into two subgroups. Type I: hereditary predisposition to colorectal cancer, a young age of onset, and carcinoma observed in the proximal colon. Type II: patients have an increased risk for cancers in certain tissues such as the uterus, ovary, breast, stomach, small intestine, skin, and larynx in addition to the colon. Diagnosis of classical HNPCC is based on the Amsterdam criteria: 3 or more relatives affected by colorectal cancer, one a first degree relative of the other two; 2 or more generation affected; 1 or more colorectal cancers presenting before 50 years of age; exclusion of hereditary polyposis syndromes. The term "suspected HNPCC" or "incomplete HNPCC" can be used to describe families who do not or only partially fulfill the Amsterdam criteria, but in whom a genetic basis for colon cancer is strongly suspected.,Function:Component of the post-replicative DNA mismatch repair system (MMR). Heterodimerizes with MLH1 to form MutL alpha. DNA repair is initiated by MutS alpha (MSH2-MSH6) or MutS beta (MSH2-MSH6) binding to a dsDNA mismatch, then MutL alpha is recruited to the heteroduplex. Assembly of the MutL-MutS-heteroduplex ternary complex in presence of RFC and PCNA is sufficient to activate endonuclease activity of PMS2. It introduces single-strand breaks near the mismatch and thus generates new entry points for the exonuclease EXO1 to degrade the strand containing the mismatch. DNA methylation would prevent cleavage and therefore assure that only the newly mutated DNA strand is going to be corrected. MulL alpha (MLH1-PMS2) interacts physically with the clamp loader subunits of DNA polymerase III, suggesting that it may play a role to recruit the DNA polymerase III to the site of the MMR. Also implicated in DNA damage signaling, a process which induces cell cycle arrest and can lead to apoptosis in case of major DNA damages.,similarity:Belongs to the DNA mismatch repair mutL/hexB family.,subunit:Heterodimer of PMS2 and MLH1 (MutL alpha). Forms a ternary complex with MutS alpha (MSH2-MSH6) or MutS beta (MSH2-MSH3). Part of the BRCA1-associated genome surveillance complex (BASC), which contains BRCA1, MSH2, MSH6, MLH1, ATM, BLM, PMS2 and the RAD50-MRE11-NBS1 protein complex. This association could be a dynamic process changing throughout the cell cycle and within subnuclear domains.,
show all
Cellular Localization:
Nuclear
show all
Tissue Expression:
Research Areas:
>>Mismatch repair ;
>>Fanconi anemia pathway
>>Fanconi anemia pathway
show all
Reference Citation({{totalcount}})
Catalog: YM6220
Size
Price
Status
Qty.
200μL
$600.00
In stock
0
100μL
$340.00
In stock
0
40μL
$190.00
In stock
0
Add to cart


Collected


Collect
Recently Viewed Products
Clear allPRODUCTS
CUSTOMIZED
ABOUT US
Toggle night Mode
{{pinfoXq.title || ''}}
Catalog: {{pinfoXq.catalog || ''}}
Filter:
All
{{item.name}}
{{pinfo.title}}
-{{pinfo.catalog}}
Main Information
Target
{{pinfo.target}}
Reactivity
{{pinfo.react}}
Applications
{{pinfo.applicat}}
Conjugate/Modification
{{pinfo.coupling}}/{{pinfo.modific}}
MW (kDa)
{{pinfo.mwcalc}}
Host Species
{{pinfo.hostspec}}
Isotype
{{pinfo.isotype}}
Product {{index}}/{{pcount}}
Prev
Next
{{pvTitle}}
Scroll wheel zooms the picture
{{pvDescr}}